This page contains information about how to access the iSDAsoil data hosted on the AWS Registry of Open Data, as well as background information about the dataset that will help you to use it. To visually explore the maps that we’ve created, please visit the iSDAsoil homepage. You can find more detailed information on how the maps were made on our technical information and further information on our FAQ pages.
Data Overview
All data has been prepared as cloud-optimised geoTIFFs (essentially geographically aware images). A single file covers the whole of Africa. Related data have been grouped together as bands in a single file. For most soil property predictions, this means that each file has 4 bands – the predicted mean value, and model error (expressed as 1 standard deviation) at 2 soil depths: 0-20cm and 20-50cm. For covariate data the number of bands is variable.
Data Access
All iSDAsoil data can be found in a publicly accessible Amazon S3 bucket at: https://isdasoil.s3.amazonaws.com/. All files have associated metadata listed according to the STAC specification. We have also included a STAC browser to allow exploration of the data in a more intuitive way. This can be found at: https://isdasoil.s3.amazonaws.com/index.html.
The STAC browser will give users the URL to download each geoTIFF, by clicking through to a specific item and navigating to the “Assets” tab. Here you will find download links for the metadata json file, and also the geoTIFF file itself. See below for an example for extractable phosphorous.
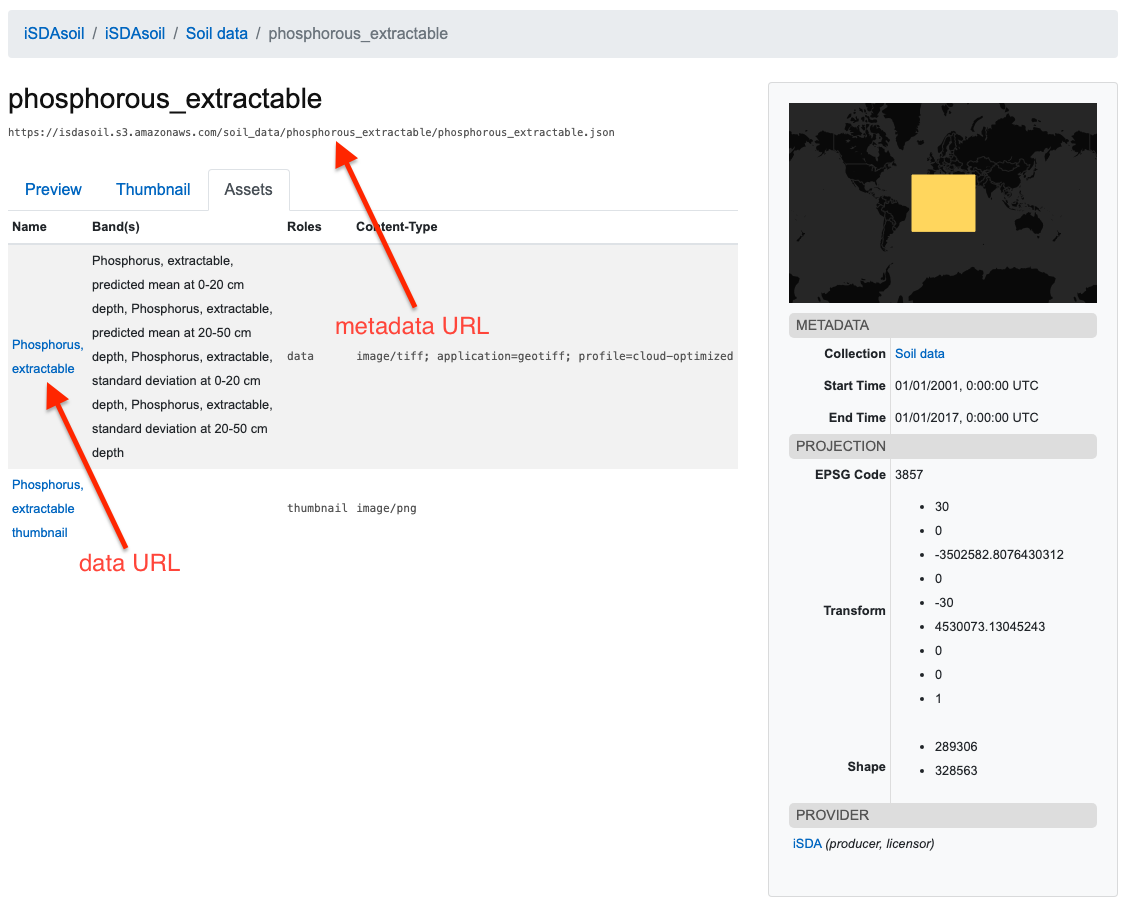
It is important to note that much of the soil data will require back-transformation of the values. Details of any back-transformation, if required, is contained within the metadata file.
Because the files are large, it may not be the best approach to attempt to download the entire file. See below for some solutions.
Viewing the data in QGIS
Due to the properties of the cloud optimised geoTIFF, It is possible to view the data in QGIS without downloading it. While this is functionally equivalent to viewing the data on the iSDAsoil homepage, the covariates used in mapping can also be viewed in this way. First the URL of the file needs to be obtained from the S3 bucket, as outlined above. Then it’s as simple as adding the layer by clicking (in QGIS):
Layer -> Add Layer -> Add raster layer -> Protocol: HTTP(S), cloud, etc.
Then add the URL of the file in the URI field. For example, for total carbon, it would be:
https://isdasoil.s3.amazonaws.com/soil_data/carbon_total/carbon_total.tif
Because most files contain multiple bands, each band will need to be viewed separately. Although analysis of the data is possible from within QGIS, we recommend using Python for any computationally intensive analysis. See below for a tutorial.
Python tutorial
We have written an iPython notebook tutorial on how to access the iSDAsoil data in Python, which walks through data access, downloading a subset and analysis. It is strongly recommended to follow this tutorial in order to gain a better understanding of the data – visit the GitHub repository at https://github.com/iSDA-Africa/isdasoil-tutorial.
Downloading data for a country or region
Perhaps the easiest way to access a subset of the data for a specific region is to use GDAL at the command line to directly download the data from s3. All you need is the URL of the file and the coordinates (in lat/lon) that specify the extent of the required bounding box. The command is structured as follows:
gdal_translate /vsicurl/{source_URL} {destination_filename} -projwin {min_lon} {max_lat} {max_lon} {min_lat} -projwin_srs EPSG:4326
So if we wanted to download the pH layer for Ghana to a file named ghana_ph.tif the command would be as follows:
gdal_translate /vsicurl/https://isdasoil.s3.amazonaws.com/soil_data/ph/ph.tif ghana_ph.tif -projwin -3.24437008301 11.0983409693 1.0601216976 4.7104621443 -projwin_srs EPSG:4326
Download the associated metadata with wget at the command line (noting that the geographical information will no longer be relevant if a subset has been downloaded):
wget https://isdasoil.s3.amazonaws.com/soil_data/ph/ph.json -O ph_metadata.json
